Home - Research activities
The main research interest of the Molecular Carcinogenesis Group (MCG) is focused on deciphering the molecular mechanisms involved in the development of neoplastic disease.
The MCG has long been involved in the investigation of the mechanisms leading to deregulation of the cell cycle and the emergence of genomic instability in cancer [see achievements i-iv]. Aberrations in the pRb/E2F, DNA damage response and repair (DDR-R) and p53 pathways/networks have been the aims of intense research. Based on the extensive molecular pathology experience that the group possesses, notable clinic-pathological observations are functionally recapitulated in vitro and in vivo, aiming to decode the underlying mechanisms driving aberrant cellular behavior. Within this context we are particularly interested in the role of cellular senescence in cancer, and other age-related pathologies, as well [achievement v].
MCG is also investigating the role of inflammation during neoplastic as well as non-neoplastic diseases. We have contributed in unraveling the systemic impact of DNA damage response aberrations in inflammatory-driven cancer [achievement vi].
In the advent of digitized medicine where the size of data exponentially increases, machine learning frameworks are entering the scene allowing us to effectively utilize the massive amount of information generated on a daily basis. Consequently, we are developing digital pathology tools that increase the efficiency and speed of diagnostic and therapeutic approaches and thus deliver personalized care opportunities for patients. Moreover we design machine learning workflow to predict drug response and survival of cancer patients [achievement viii].
Within the frame of physiological and pathological human development we are interested also in human evolution. As a result and in collaboration with Professor Katerina Harvati, from the University of Tübingen, we published in Nature a series of important findings in the field of palaeoanthropology that are related to the early dispersal and presence of modern humans in Eurasia. Moreover, in a bone fragment adjacent to the human the skulls found we identified, using an innovative reagent that we developed [SenTraGor - see achievement v], the presence of lipofuscin a potent biomarker of senescence (Pharmacology –Therapeutics 2018). This finding depicts how resilient is lipofuscin to time and cellular degradation processes [achievement vii].
MCG has recently expanded its research interests towards Nanotechnology applications in human disease and life sciences. In collaboration with the Laboratory of General Chemistry at the National Technical University of Athens (N.T.U.A.), we aim to develop innovative nanoparticles with the potential to act as anticancer agents, in parallel with the discovery of advanced drug delivery systems for targeted therapeutic approaches.
Apart from the research activities, MCG is providing molecular pathology services. As an active collaborator of the European Society of Pathology, MCG is participating in the annually K-ras European Quality Assurance Program, related to the anti-EGFR therapy.
Major Achievements
Our group played a major role in:
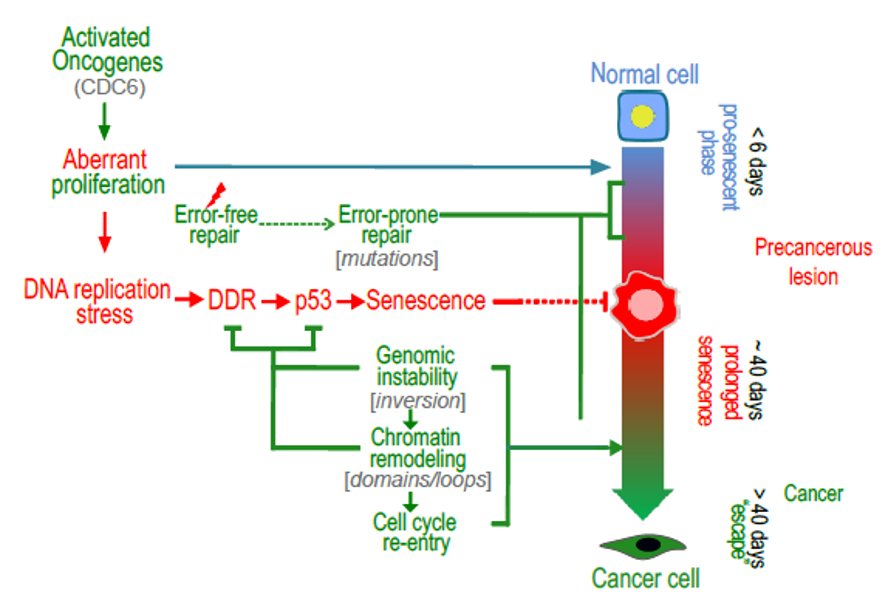
i) Establishing the “Oncogene-induced DNA damage model for cancer development” (Science 2008; Nat Rev Mol Cell Biol 2010; J Pathol 2018) (Figures 1-6, 8, 13, 14). Based on: Cancer Res 2001; J Pathol 2002; J Pathol 2004; Nature 2005; Nature 2006; J Pathol 2006; Cancer Res 2007 (1); Oncogene 2008; Blood 2008; Am J Pathol 2009; Nat Genet 2011; Nat Cell Biol 2011; Cancer Res 2012; Cell Death Differ 2014; Cell Mol Life Sci 2014; Cell Rep 2015; Nat Cell Biol 2016; Genomics 2018; Genome Biol 2018; Mol Cell 2021; EMBO Rep 2021 and currently the broader role of DNA Damage response pathway in disease development (Cell 2016).
ii) Demonstrating that oncogene-induced senescence is a DNA damage stress response acting as a barrier to cancer (Nature 2006; Cancer Res 2007; Am J Pathol 2009; Curr Opin Cell Biol 2010; Nat Cell Biol 2011; Cell Death Differ 2015; Nat Cell Biol 2016; BMC Genomics 2018; Genome Biol 2018) (Figures 2,5,6).
iii) Clarifying the functional interplay and the timeline of events underlying the two major antitumor checkpoint responses, i.e. DDR and ARF (Nat Cell Biol 2013; Cell Death Diff 2013; Cell Cycle 2014) (Figure 2).
iv) Revealing the oncogenic role of replication licensing factors Cdc6 and Cdt1 by inducing DNA replication stress and deregulating transcription (Am J Pathol 2004; Nature 2006; Cancer Res 2007, J Cell Biol 2011; Transcription 2012; Semin Cancer Biol 2016; Nat Commun 2016; Nat Cell Biol 2016; PNAS 2016; BMC Genomics 2018; Genome Biol 2018; Mol Cell 2021) (Figures 2-6).
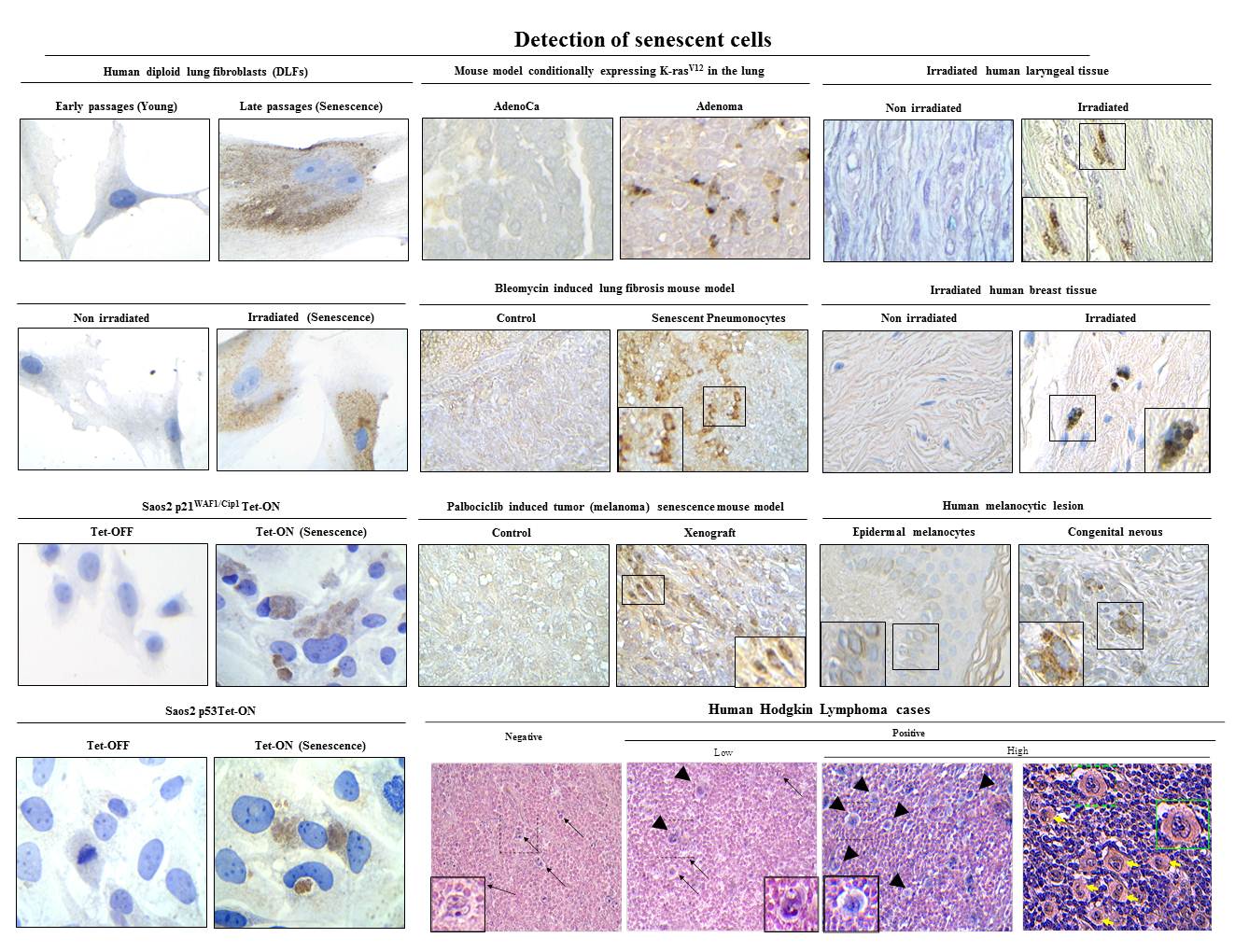
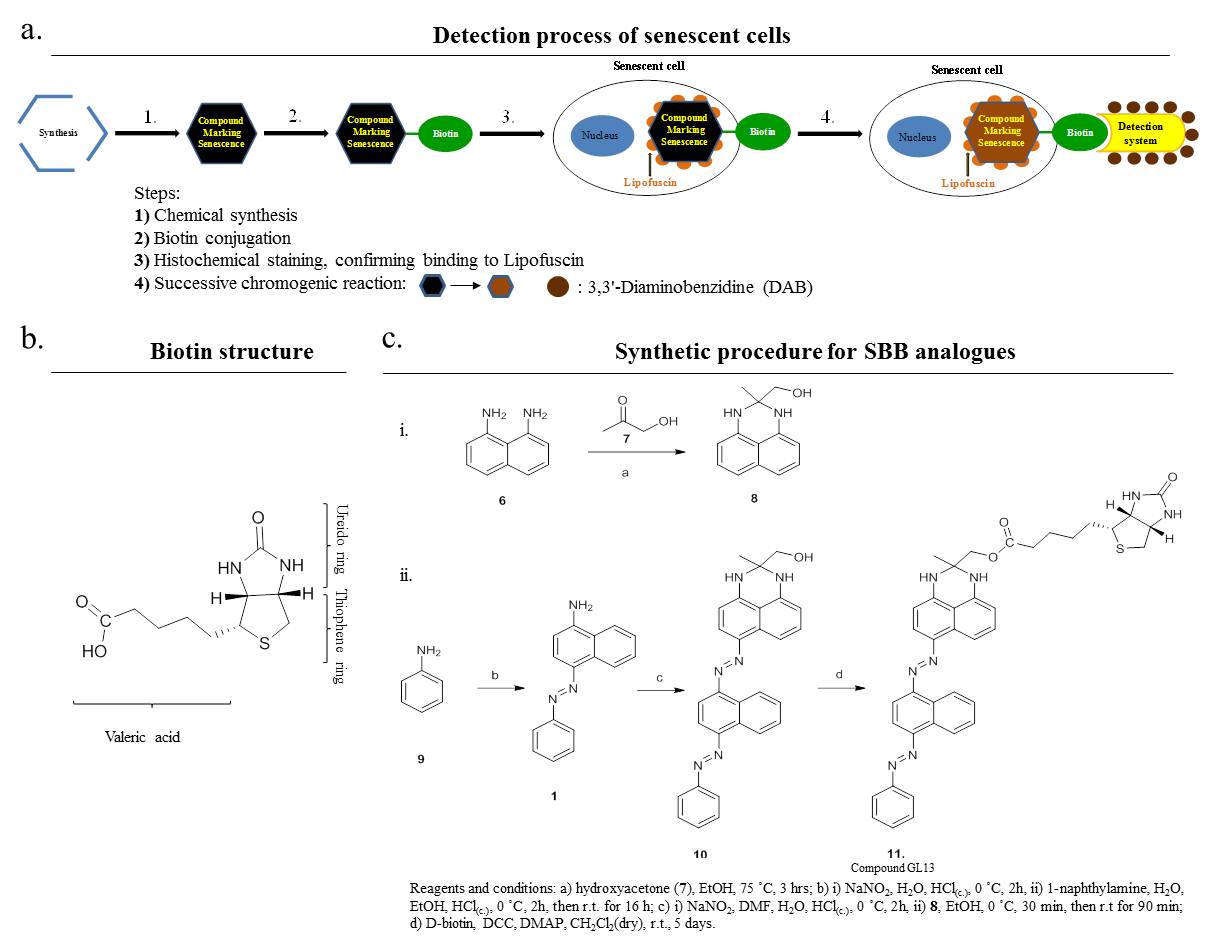
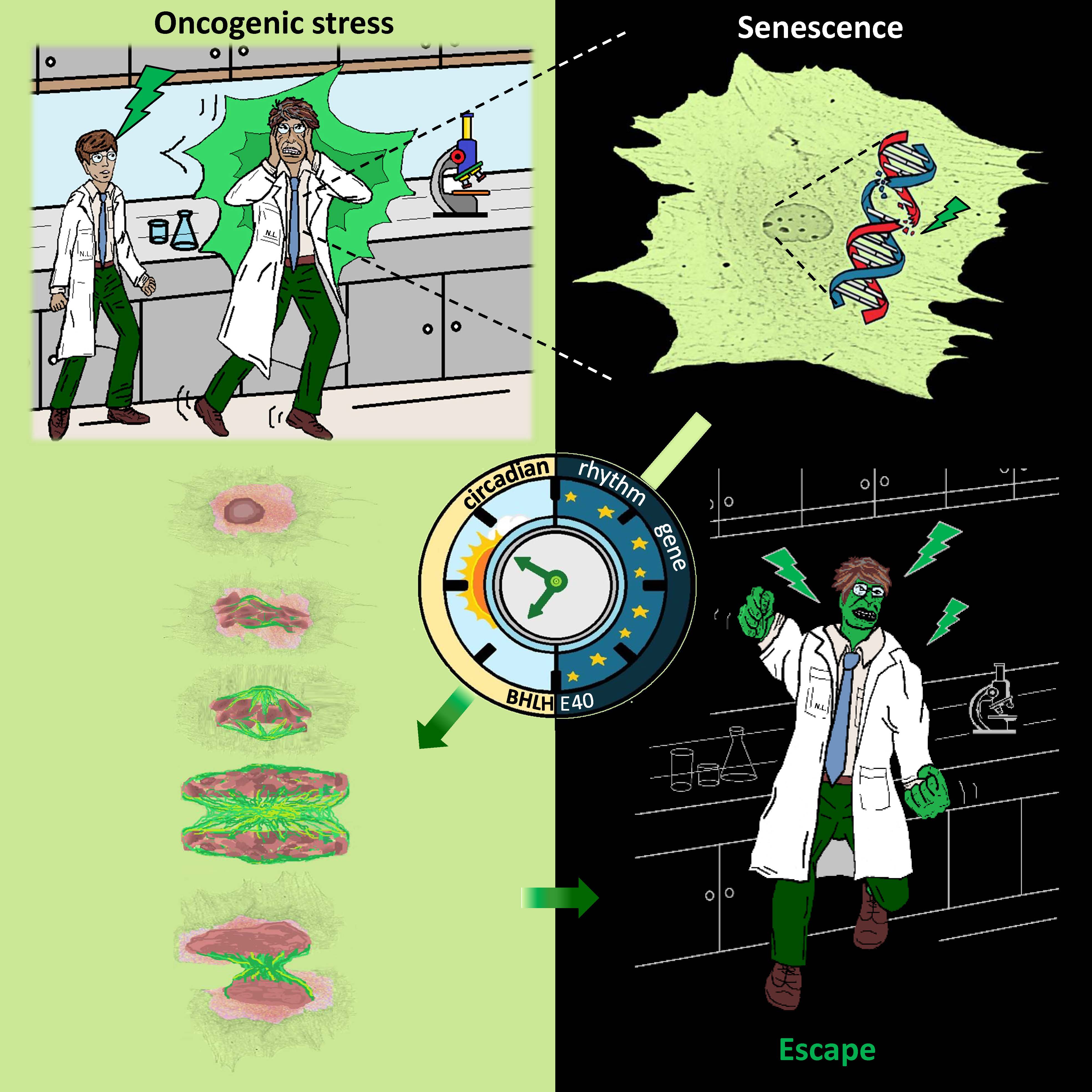
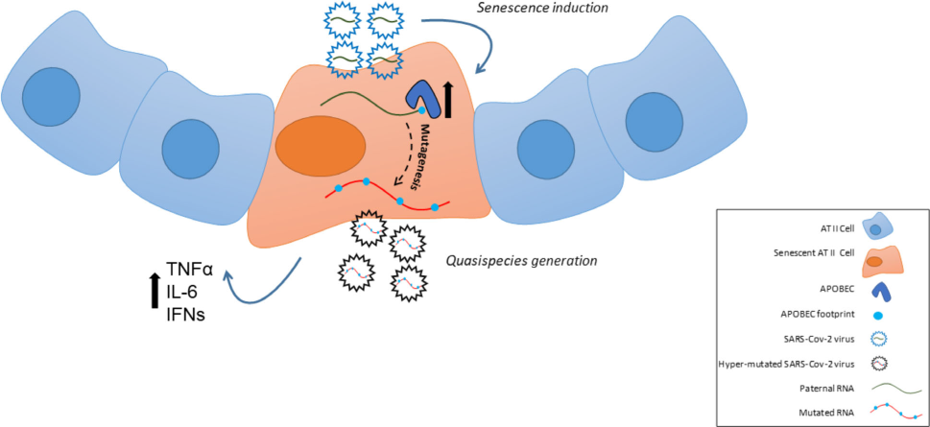
v) Understanding how cellular senescence is involved in age-related pathologies, including cancer and viral infections (EMBO J 2003; Lab Invest 2005; Nature 2006; Am J Pathol 2009; Aging 2013; Aging Cell 2013; Stem Cells 2013; Cell Death Differ 2015; Mech Ageing Dev 2016; Aging Cell 2017; J Pathol 2018; Int J Mol Sci 2018; Pharmacol Ther 2019 (1); Nat Commun 2020 (1); Nat Commun 2020 (2); Mol Cell 2021; Europ Res J 2022; Physiol Rev 2022) (Figures 2,5,6,8,10,13,15). Our research has led to the development of a pioneer senescence biomarker (commercially available under the trademark SenTraGorTM) able to detect senescence in any biological material including archival (Aging 2013; Aging Cell 2017; Int J Mol Sci 2018; Meth Mol Biol 2019; Pharmacol Ther 2019 (1); Cell 2019; Nat Protoc 2021) (Figures 10,11).
vi) Contributing to our understanding of the role that inflammation plays in cancer development (Cancer Res 2007 (2); Cancer Cell 2013; Cancer Cell 2014; Pharmacol Ther 2015; Cell Death Differ 2015; Cell Rep 2016; Nat Commun 2018; Nat Commun 2020 (2)) (Figures 7,8).
vii) Providing evolutionary evidence that modern humans dispersed into Europe and Asia (150 thousand years) earlier than initially thought (Nature 2019; Sci Adv 2022) (Figure 9) (and recently supported by ERC AdG 2020).
viii) Development of machine learning algorithms to predict drug response and survival of cancer patients (Pharmacol Ther 2019 (2); Cell Reports 2019; Cancer Res 2020) (Figure 12).
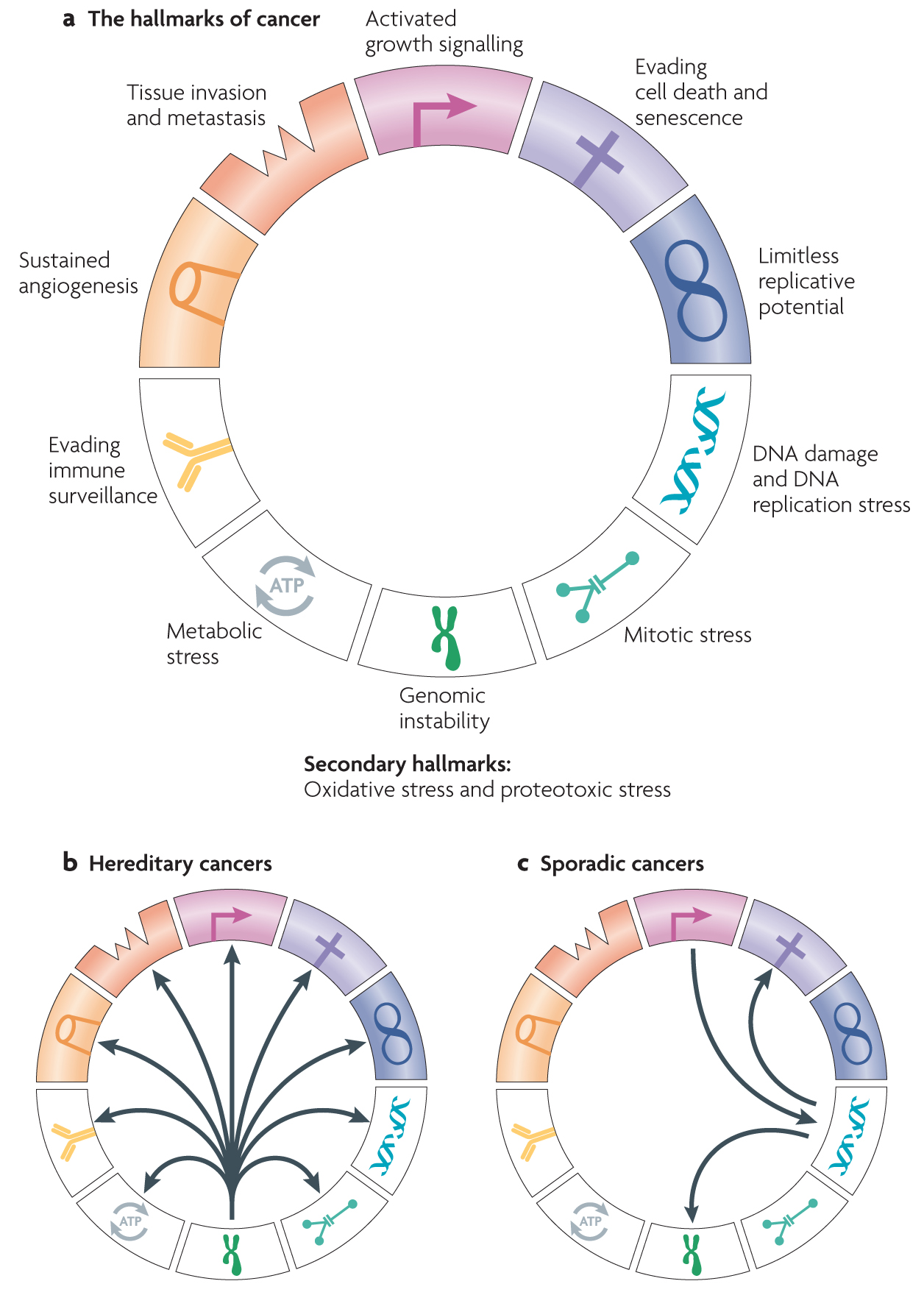
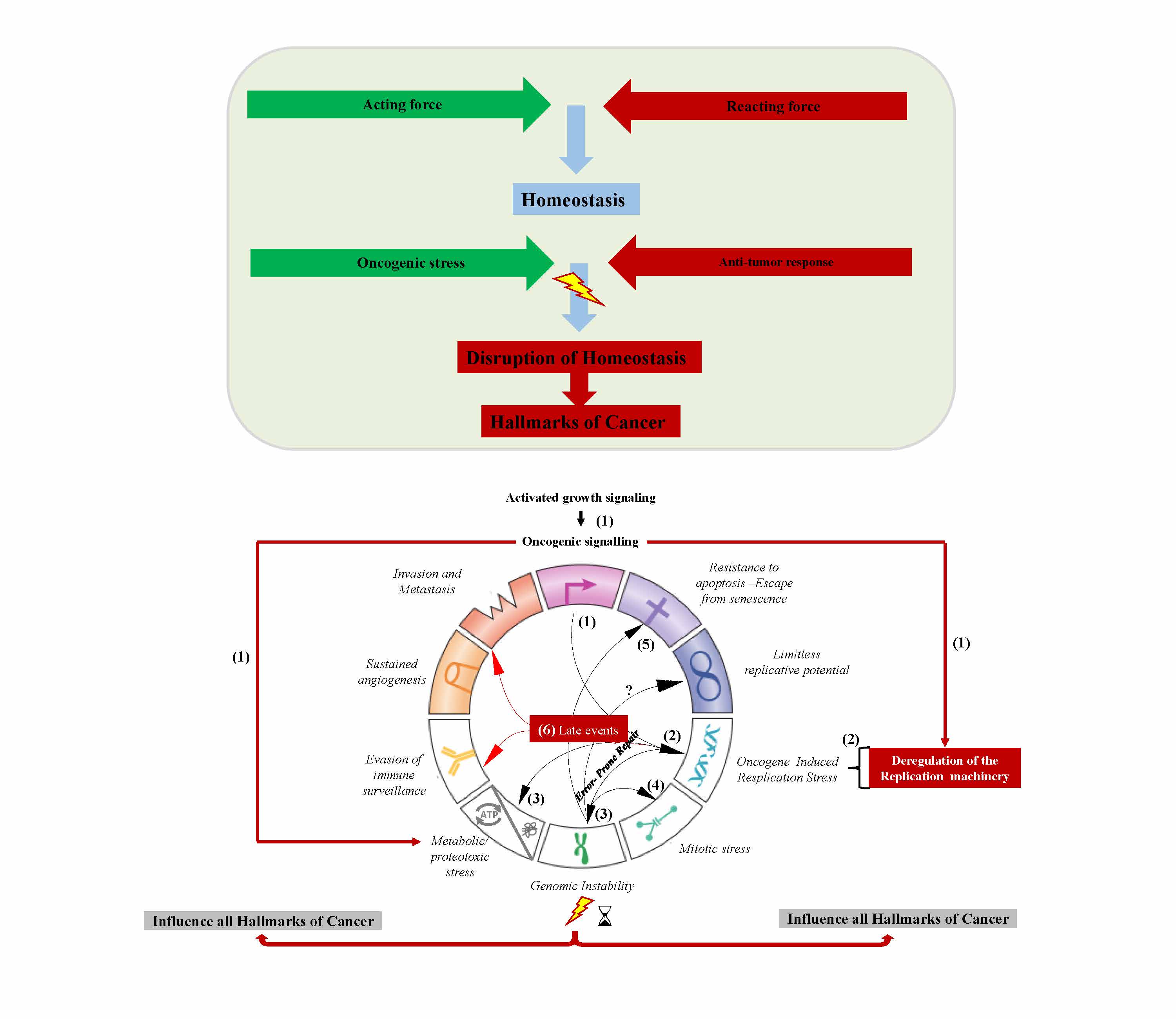
1. Genomic instability — an evolving hallmark of cancer
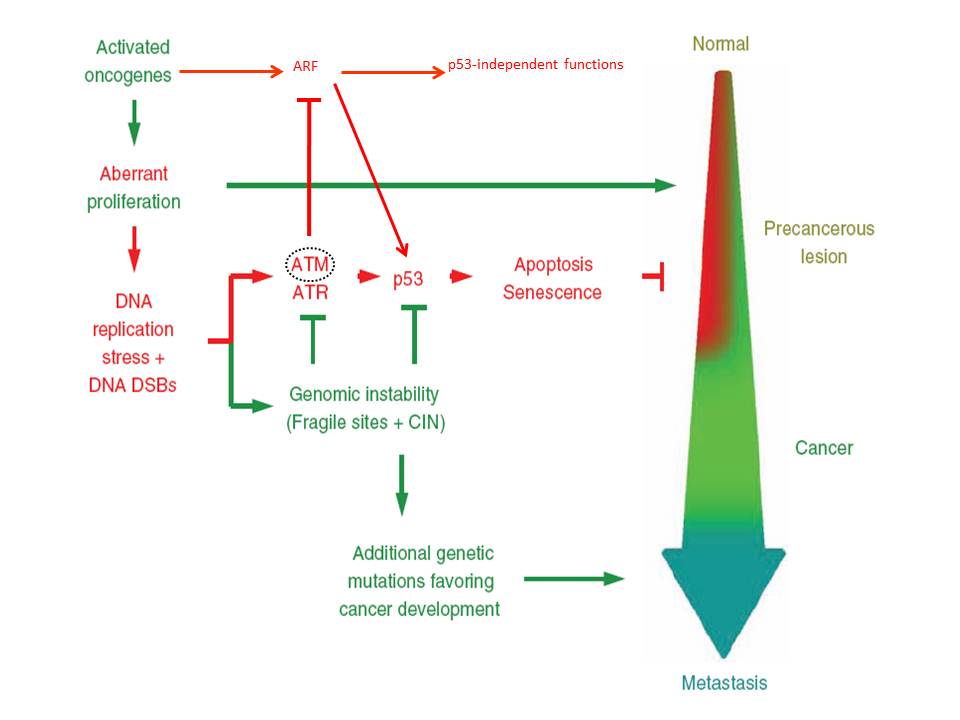
2. An Oncogene-Induced DNA Damage Model for Cancer Development
Science 2008
Nature 2005
Nature 2006
Nature Cell Biol 2013
CDD 2013
3. The role of Cdc6 in cancer progression

Am J Pathol 2004
Cancer Res 2007
J Cell Biol 2011
Transcription 2012
4. Oncogenic Cdc6 as a molecular switch during cancer development
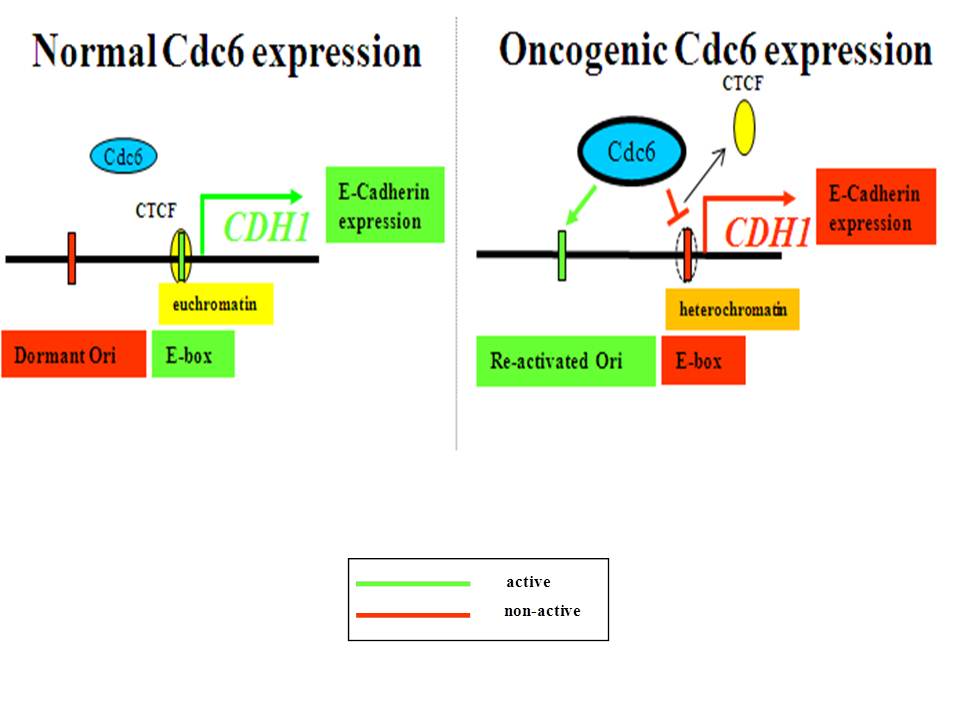
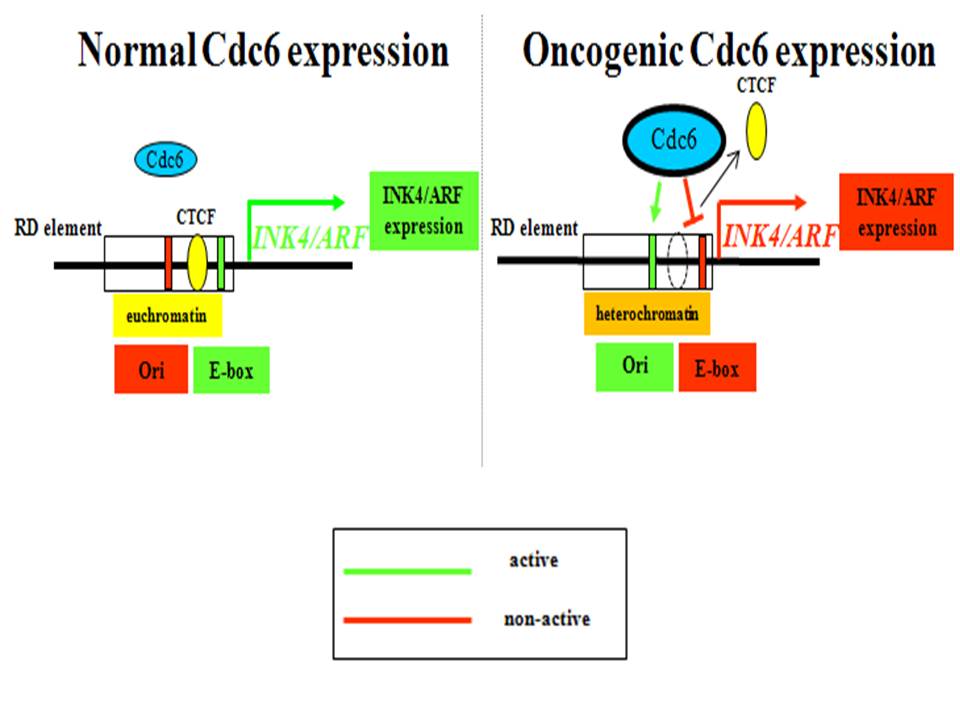
J Cell Biol 2011
Transcription 2012
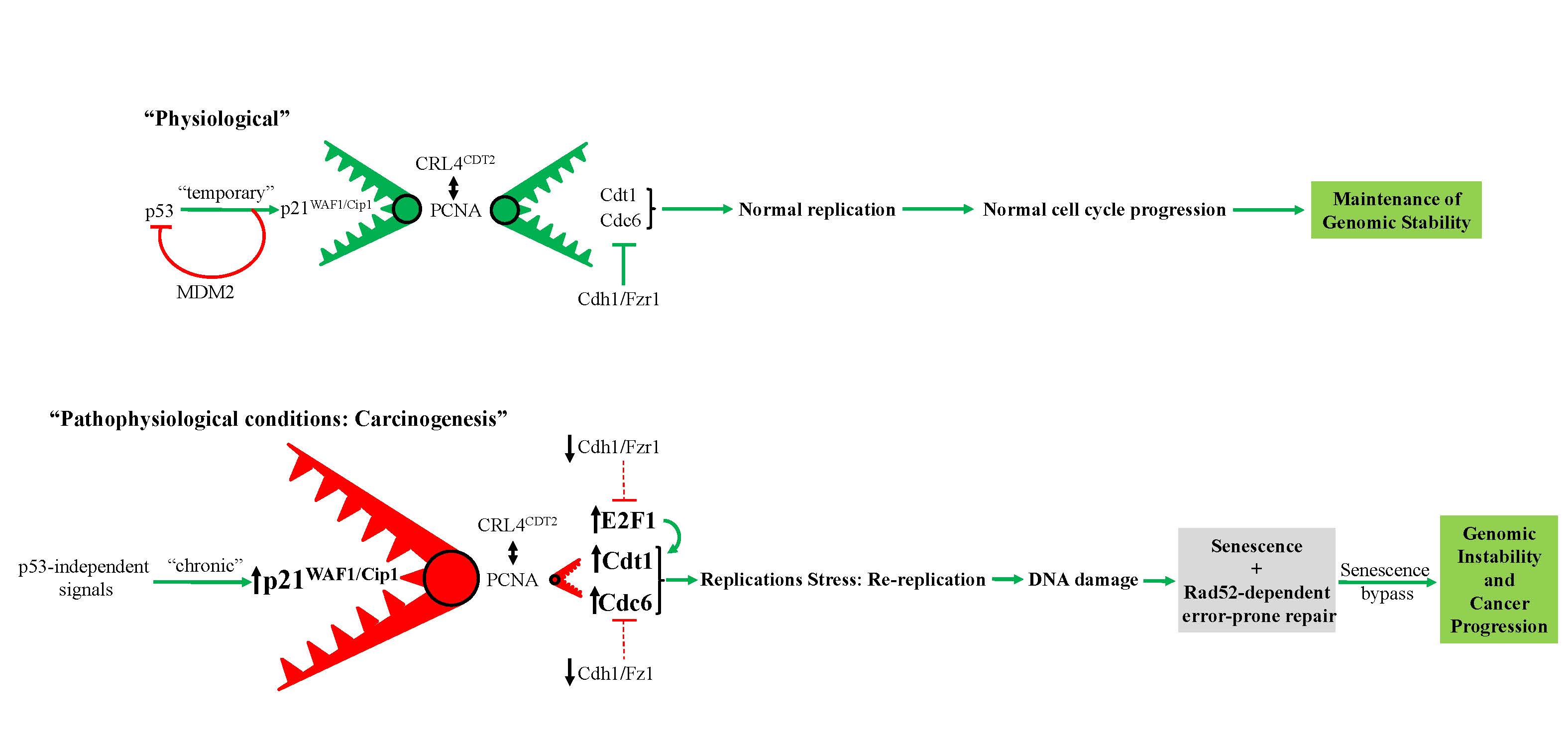
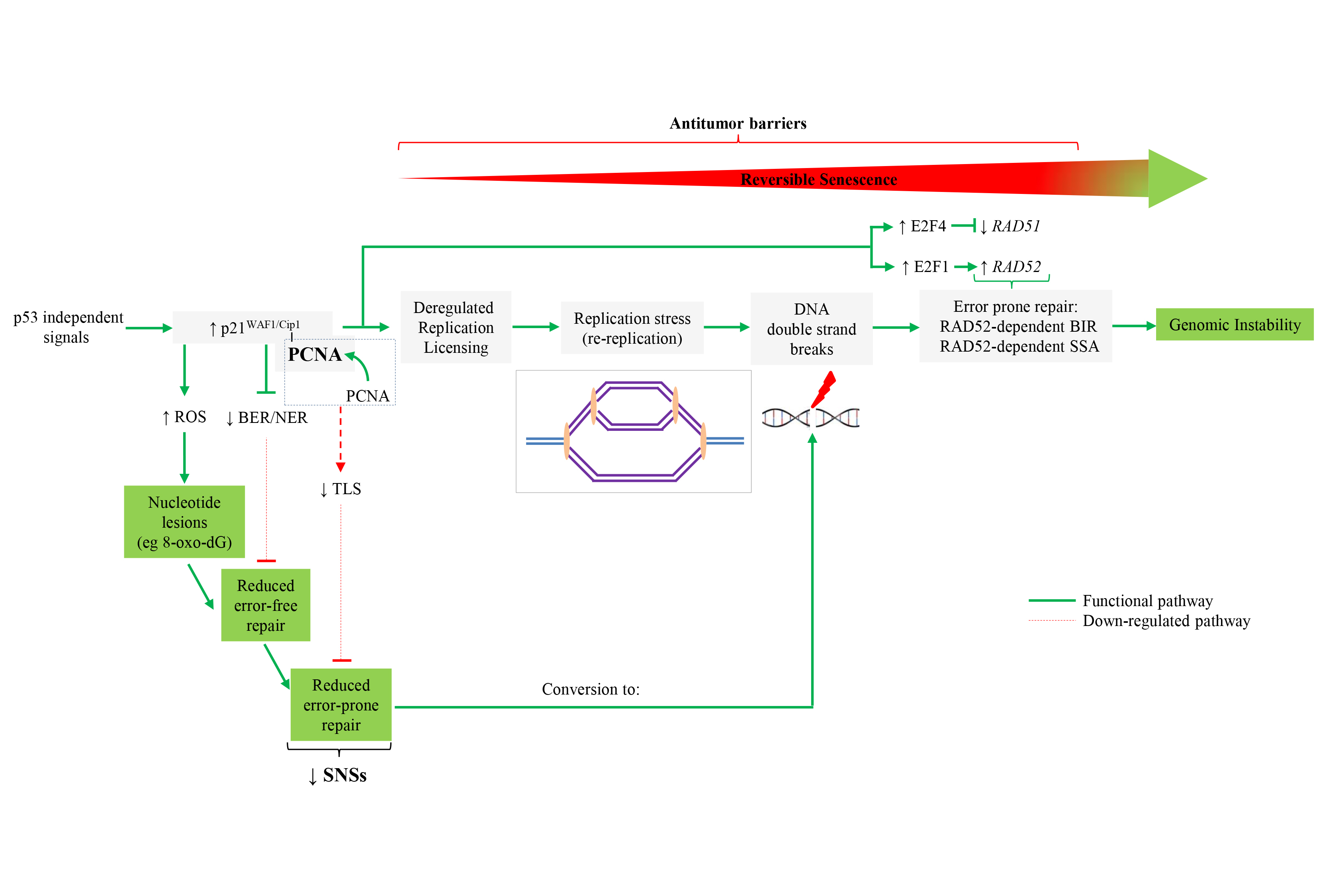
5. Prolonged expression of p21WAF1/Cip1 in p53-null cells as a driving force for cancer progression
Re-replicating cell (video)
Escaped and dividing cells (video)
RAD52 recruitment OFF cells (video)
RAD52 recruitment ON cells (video)
6. SenTraGor: a novel reagent to detect senescent cells
Aging 2013
Meth Mod Biol 2017
Aging Cell 2017
Meth Mol Biol 2018
Cell 2019
7. The non cell-autonomous role of mutant p53 gain-of-function: reprogramming the microenvironment

i)Cancer Cell & Microenvironment 2014
iia)Nat Commun 2018
iib)Nat Commun 2018
8. Integrating the DNA damage and protein stress responses during cancer development and treatment
9. Apidima Cave fossils provide earliest evidence of Homo sapiens in Eurasia
a.
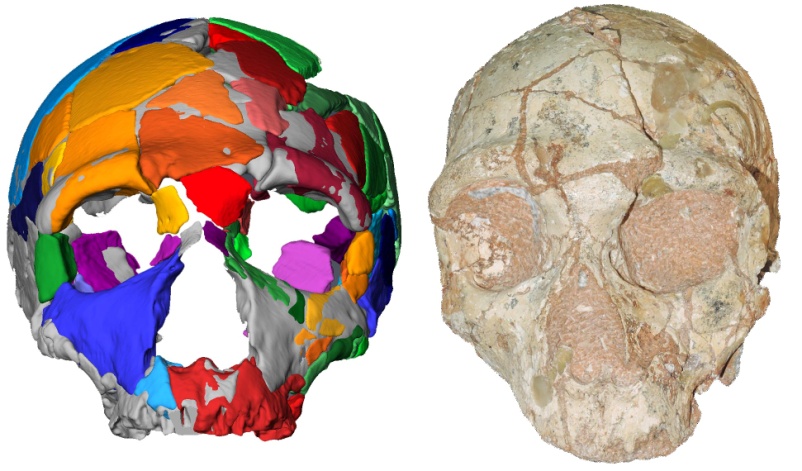
(a) Analysis of two human fossil skulls (Apidima 1 and Apidima2) found during excavations at Apidima cave in Mani, Peloponnese, showed that fossil find Apidima 1 was dated about 210.000 years old. It represented a modern human with archaic characteristics, indicating an early Homo sapiens. Apidima 2 was dated 170.000 years old, with Neanderthal features. These findings imply that early modern humans spread into Eurasia 150 thousand years earlier than thought.
b.
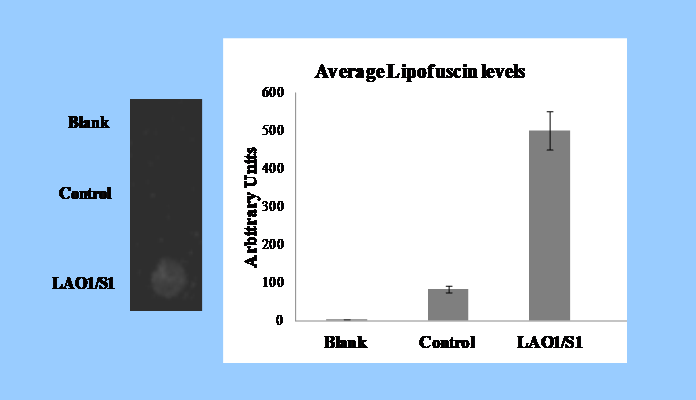
(b) In a bone fragment (LAO1/S1) found adjacent to the human skull Apidima 1, we identified, using an innovative reagent (SenTraGor) that we developed, the presence of lipofuscin a potent biomarker of senescence.
Nature 2019
Pharmacology –Therapeutics 2018
Cell 2019
10. Cellular Senescence: Defining a Path Forward
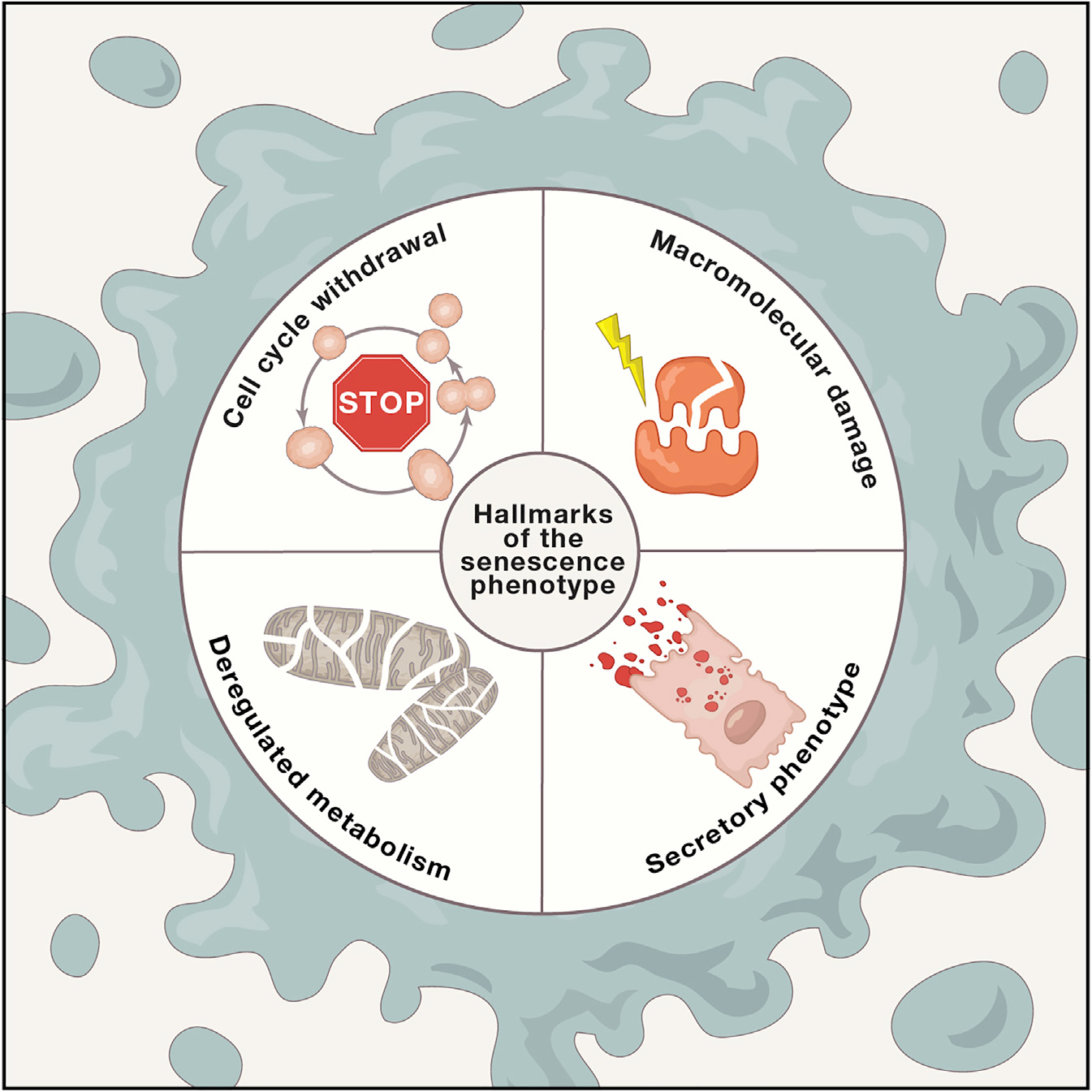
The Hallmarks of the Senescence Phenotype
SeneQuest: Free web-resource application for identification of factors and conditions involved in senescence
11. Algorithmic assessment of cellular senescence in experimental and clinical samples
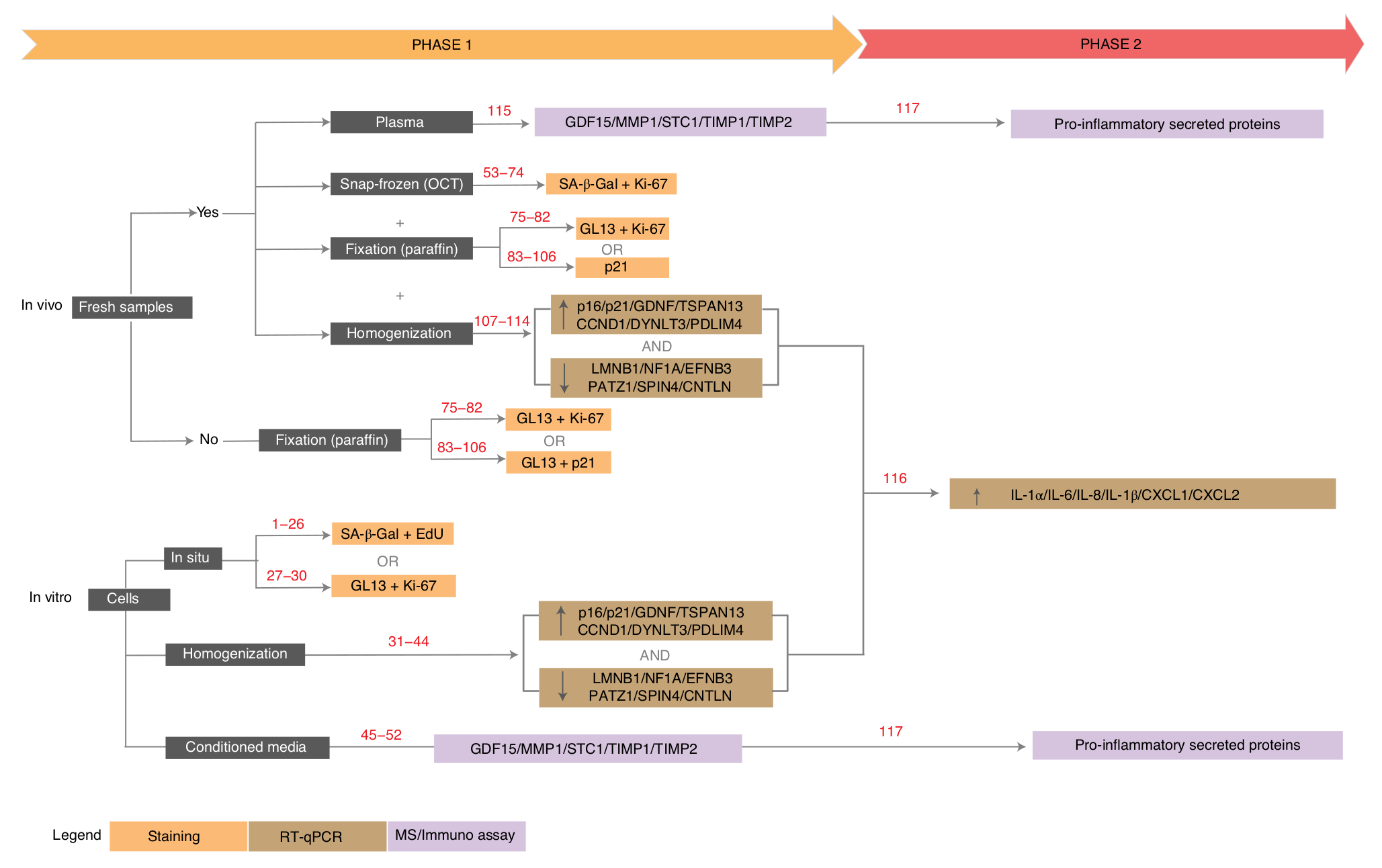
12. Machine learning algorithms to predict drug responses in cancer
A.
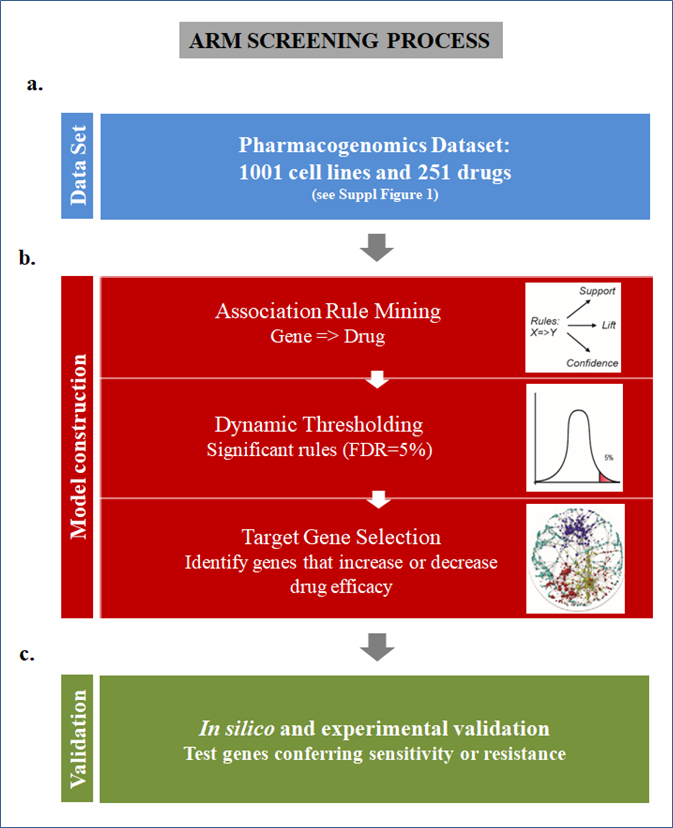
Rshiny: Free web-resource application connecting molecular defects with drug response
Pharmacology – Therapeutics 2019
B.
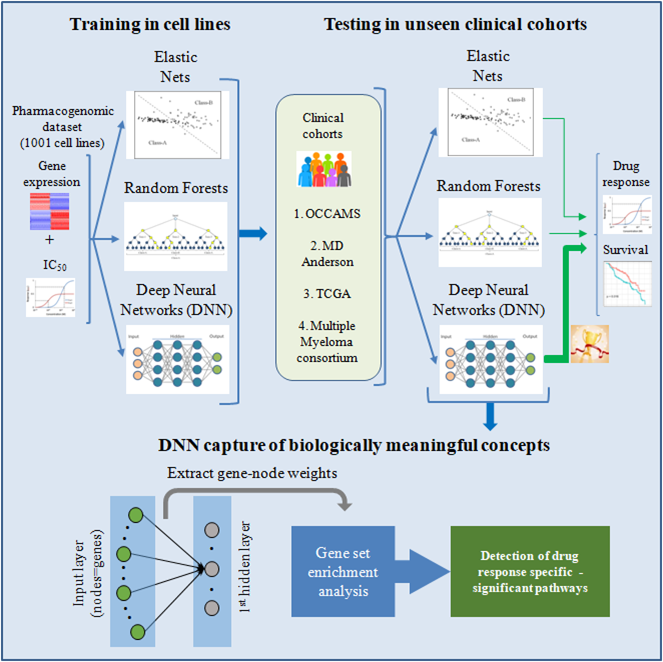
14. RASSF1A-mediated mechanism controlling tumor dedifferentiation and aggressive oncogenic behavior
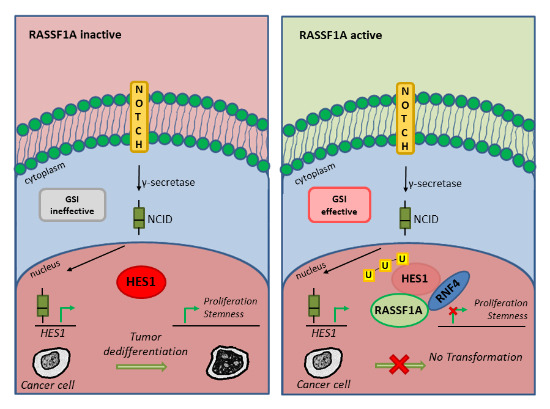
FEBS 2021
Europ Res J 2022
Europ Res J 2022 (commentary)
Europ Res J 2022 (editorial)
16. Machine learning: a tool to shape the future of medicine

Handbook of Machine Learning Applications for Genomics
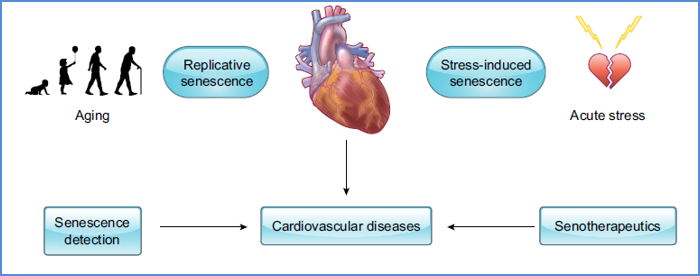
17. Cellular senescence and cardiovascular diseases
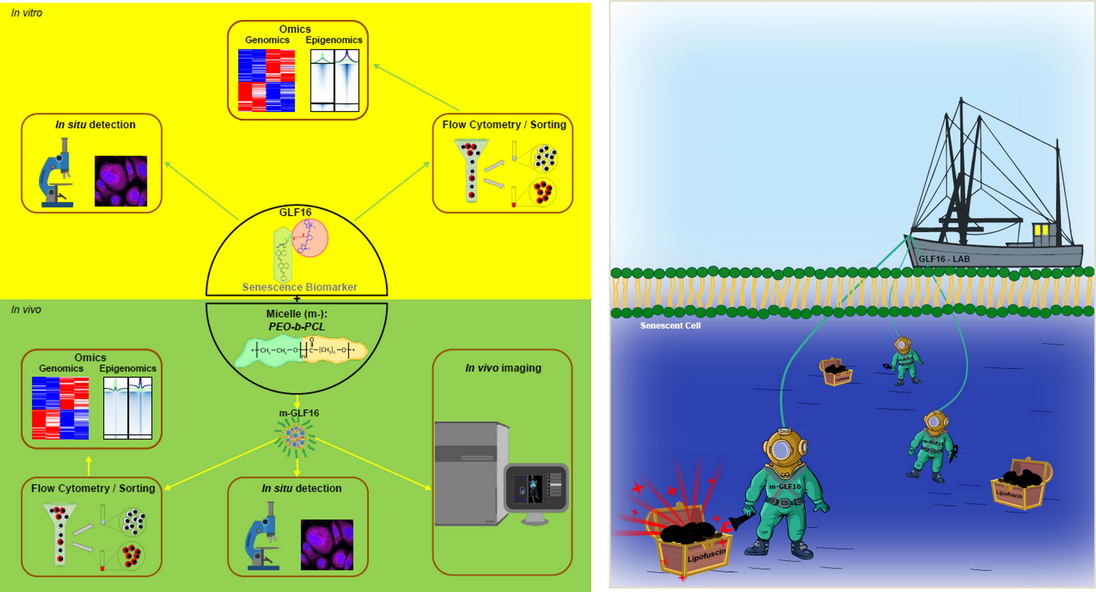
19. Implication of senescence in giant cell arteritis
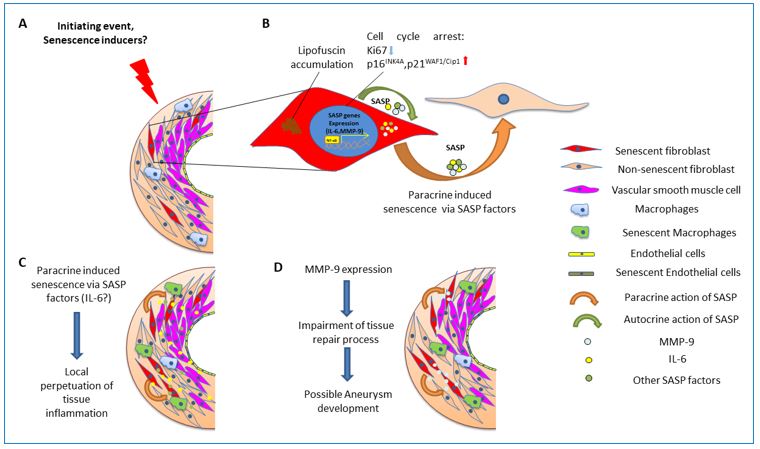
|
|
|
Prof. Vassilis G. Gorgoulis
Laboratory of Histology-Embryology
Chair of Clinical Molecular Pathology, Ninewells Hospital and School of Medicine
University of Dundee, Dundee, UK
Biomedical Research Foundation of the Academy of Athens
Faculty Institute for Cancer Sciences, University of Manchester, Manchester Centre for Cellular Metabolism,
EMBO member
European Academy
Academia Europaea member
Intelligencia.ai, 180 Varick Street, 6th Floor, New York, NY 10014, USA
Office Tel: 0030 210-7462352 |
News
Error: No articles to display